Abstract
Background: Enterobacteriaceae that produce extended-spectrum β-lactamase (ESBL) have emerged as a serious threat, with variable rates depending on geographic region. We determined the prevalence of ESBL-producing Escherichia coli, Klebsiella pneumoniae, K. oxytoca and Proteus mirabilis in bloodstream infections in Toronto from 2006 through 2016.
Methods: All patients with E. coli, K. pneumoniae, K. oxytoca and P. mirabilis isolated from blood in a tertiary care microbiology laboratory in Toronto between 2006 and 2016 (1 isolate per species per patient per year) were included in this retrospective cohort study. Organisms were identified by conventional methods, and susceptibility testing was performed according to Clinical and Laboratory Standards Institute standards. Screening for ESBL and phenotypic confirmatory testing were done with a modified Clinical and Laboratory Standards Institute method. ST131 clonal type was determined by means of an established protocol.
Results: The proportion of ESBL-producing E. coli isolates increased significantly between 2006 and 2016, from 6.4% (19/296) to 17.3% (89/513) (p < 0.001). This trend was seen in both intensive care units and emergency departments. Concurrently, the proportion of ST131 among ESBL-producing E. coli also increased significantly, from 31.6% (6/19) in 2006 to 73.0% (65/89) in 2016 (p = 0.03). Among ESBL-producing E. coli, significant resistance was noted to multiple antimicrobial classes. Comparable increases in the proportion of ESBL-producing K. pneumoniae, K. oxytoca and P. mirabilis were not noted.
Interpretation: We observed a significant increase in the proportion of ESBL-producing E. coli in bloodstream infections in Toronto temporally correlated with an increase in the ST131 clonal type. Recognition of this dramatic rise is important to inform empiric antibiotic treatment.
The increase in antimicrobial-resistant organisms has reached alarming proportions. The Public Health Agency of Canada has stated that antimicrobial resistance is an increasingly serious threat to global public health.1 The World Health Organization identified 12 bacterial groups that are of great concern.2 The Centers for Disease Control and Prevention identified 18 microorganisms that pose urgent, serious and concerning threats.3 Enterobacteriaceae that produce extended-spectrum β-lactamase (ESBL) are included on their lists as a “Priority 1: Critical” organism2 or a “serious threat.”3 Within the Enterobacteriaceae, ESBL-producing Escherichia coli is a frequent cause of infection, with bloodstream infections being associated with substantial mortality rates (14%–30%).4–6 Extended-spectrum β-lactamases are enzymes that are capable of hydrolyzing most β-lactam antimicrobials, and the plasmids on which they are transported frequently contain other resistance genes.7,8 This leaves few treatments available to physicians and increases the likelihood of E. coli’s developing resistance to all available drugs.
The prevalence of ESBL production among E. coli isolates has increased worldwide and varies between regions. Rates as high as 70% have been reported in regions in Southeast Asia.9,10 There is a paucity of recent published Canadian data. Historically, the prevalence of ESBL production in E. coli in Canada was less than 5% until the last decade.11 Peirano and colleagues12 reported a rate of 14% among bloodstream E. coli isolates in 2010, and a national rate of 7.1% among E. coli from blood, urine, respiratory and wound specimens combined collected from 2007 to 2011 was reported as part of the CANWARD national surveillance study.11 In both studies, these increases correlated with an increased prevalence of the ST131 clone of E. coli, which is thought to be a driving force behind the rise in ESBLs worldwide.7,13 Identifying increases in ESBL prevalence is important to inform empiric treatment, as inappropriate therapy has been linked to worse outcomes.14–16 The purpose of our study was to determine the local prevalence of ESBL-producing E. coli, Klebsiella pneumoniae, K. oxytoca and Proteus mirabilis among blood culture isolates over the last 11 years in the Toronto region.
Methods
Design and setting
This retrospective cohort study was performed through the University Health Network/Sinai Health System Department of Microbiology’s large tertiary care academic microbiology laboratory, which serves 4 acute care hospital sites, 4 rehabilitation centres and 3 long-term care/palliative/mental health sites in Toronto. With the exception of a neonatal intensive care unit, the population served is an adult population.
Study participants and bacterial isolates
All patients with bacteremia due to E. coli, K. pneumoniae, K. oxytoca or P. mirabilis between Jan. 1, 2006, and Dec. 31, 2016 at the University Health Network and Sinai Health System were included in this study. The condition of all patients contributing blood culture isolates remained stable over the 11-year period. We reviewed the records of all patients with E. coli, K. pneumoniae, K. oxytoca or P. mirabilis isolates recovered from blood cultures in the laboratory between 2006 and 2016 as identified in the laboratory information system. Following standard Clinical and Laboratory Standards Institute recommendations for annual antibiograms so that data are representative of typical antibiograms,17 we included 1 isolate per species per patient per year. Organisms isolated between January 2006 and March 2013 were identified with the Vitek 2 system (bioMérieux), and organisms isolated from April 2013 to December 2016 were identified with the Vitek MS MALDI-TOF (matrix-assisted laser desorption ionization — time of flight) mass spectrometry system (bioMérieux).
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing was performed at the time of isolation with the Vitek 2 system or disk diffusion (Oxoid) according to Clinical and Laboratory Standards Institute standards current at the time of testing.18 All isolates were tested for susceptibility to ciprofloxacin, trimethoprim–sulfamethoxazole, tobramycin, gentamicin, and either meropenem or imipenem. Ertapenem susceptibility results were available from 2009 to 2016 only.
Extended-spectrum β-lactamase screening and phenotypic confirmatory testing
Before October 2015, E. coli, K. pneumoniae, K. oxytoca and P. mirabilis isolates were screened for ESBL production as part of routine laboratory procedures. All isolates with a cefpodoxime minimum inhibitory concentration of 1 mg/L or greater with the Vitek 2 system were considered screen-positive. A double-disk test was used for phenotypic confirmation of all screen-positive isolates by means of a modified protocol based on the work of M’Zali and colleagues.19 Mueller–Hinton agar was inoculated with 0.5 McFarland bacterial suspension, and an amoxicillin–clavulanic acid disk was placed in the centre; cefpodoxime, ceftriaxone, ceftazidime and aztreonam disks were placed around it, with 15 mm between disk edges. After 18–24 hours of incubation, class A ESBL production was identified by potentiation of the inhibition zone around the amoxicillin–clavulanic acid disk near any of the surrounding disks. After October 2015, susceptibility results were reported based on Clinical and Laboratory Standards Institute interpretations with the Vitek 2 system. All E. coli, K. pneumoniae, K. oxytoca and P. mirabilis isolates from this period that were or were not susceptible to ceftriaxone were subcultured twice from stock stored in glycerol broth at −80°C, and ESBL production was identified retrospectively by means of the double-disk test. K. oxytoca isolates with a phenotype suggestive of K1 β-lactamase hyperproduction (positive with the double-disk test but susceptible to ceftazidime) were considered non–class-A ESBLs. Isolates that could not be recovered were excluded from the study.
ST131 detection among E. coli producing extended-spectrum β-lactamase
All E. coli isolates from 2006 through 2016 were subcultured twice from stock stored in glycerol broth at −80°C, and DNA extraction was completed with a boiled lysate protocol.20 ST131 clones were identified by means of a polymerase chain reaction assay that targeted the pabB gene based on the method previously described by Clermont and colleagues.21 Isolates that could not be recovered were excluded from the ST131 analysis.
Statistical analysis
We used the χ2 test for trends to analyze trends in susceptibility, proportion of ESBL-producing isolates and proportion of ST131 E. coli isolates over the study period. We used the Fisher exact test to analyze differences in susceptibility between ESBL-producing and non–ESBL-producing E. coli. All analyses were conducted with GraphPad InStat (Graph-Pad Software). We performed analysis for all blood culture isolates and carried out subset analyses for intensive care unit and emergency department isolates. A 2-sided p value of < 0.05 was considered statistically significant.
Ethics approval
This study did not involve patients or accessing patients’ medical records; thus, ethics review was not needed.
Results
During the study period, 6754 patients (3427 males [50.7%], 3327 females [49.3%]) met the study inclusion criteria. They had a total of 4615 E. coli, 1541 K. pneumoniae, 261 K. oxytoca and 342 P. mirabilis blood culture isolates. One E. coli isolate and 1 K. pneumoniae isolate from 2016 were excluded because of a presumed loss of the resistance plasmid detected on ESBL confirmatory testing, and 2 K. pneumoniae isolates from 2016 were excluded as they could not be recovered from the glycerol stock; this left a total of 6755 isolates that could be analyzed with regard to ESBL production. Ten ESBL-producing E. coli isolates (1 from each of 2008, 2010 and 2013, 2 from 2009 and 5 from 2011) could not be recovered from the glycerol stock and were excluded from ST131 analysis.
Extended-spectrum β-lactamase production
E. coli (542 [11.7%]) and K. pneumoniae (81 [5.3%]) isolates that produced ESBL predominated. There were 10 (3.8%) ESBL-producing K. oxytoca isolates and 3 (0.9%) ESBL-producing P. mirabilis isolates.
Trends in extended-spectrum β-lactamase production
E. coli
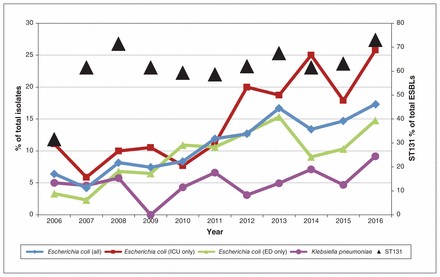
The proportion of ESBL-producing E. coli isolates increased significantly over the study period, from 6.4% (19/296) in 2006 to 17.3% (89/513) in 2016 (p < 0.001, χ2 test for trend) (Table 1, Figure 1). This trend was observed in both the intensive care unit and the emergency department, where the proportion of ESBL-producing E. coli isolates increased from 11.1% (1/9) to 25.8% (8/31) (p = 0.02, χ2 test for trend) and from 3.3% (6/180) to 14.8% (47/318) (p < 0.001, χ2 test for trend), respectively (Table 2, Figure 1). In addition, the frequency of the ST131 clonal type within ESBL-producing E. coli isolates increased from 31.6% (6/19) to 73.0% (65/89) over the study period (p = 0.03, χ2 test for trend) (Table 1, Figure 1).
Proportion of Escherichia coli, Klebsiella pneumoniae, K. oxytoca and Proteus mirabilis blood culture isolates that produced extended-spectrum β-lactamase, including proportion of ST131 clonal type within E. coli isolates that produced extended-spectrum β-lactamase, 2006–2016
Proportion of Escherichia coli and Klebsiella pneumoniae blood culture isolates that produced extended-spectrum β-lactamase (ESBL), altogether and in intensive care units (ICUs) and emergency departments (EDs), and proportion of ST131 clonal type within E. coli isolates that produced ESBL, 2006–2016.
Proportion of Escherichia coli blood culture isolates that produced extended-spectrum β-lactamase in intensive care units and emergency departments, 2006–2016
K. pneumoniae, K. oxytoca and P. mirabilis
During the study period, ESBL production among K. pneumoniae isolates showed an upward trend, from 5.0% (6/120) of isolates in 2006 to 9.2% (18/196) in 2016 (p = 0.04, χ2 test for trend). The proportion of ESBL-producing K. oxytoca isolates decreased (p = 0.002, χ2 test for trend), although the absolute number of ESBL-producing isolates remained low throughout the study period. The proportion of ESBL-producing P. mirabilis isolates remained relatively stable (Table 1). Owing to the small number of these isolates, we did not perform subset analysis of trends in the intensive care unit and the emergency department.
Antimicrobial susceptibility
E. coli
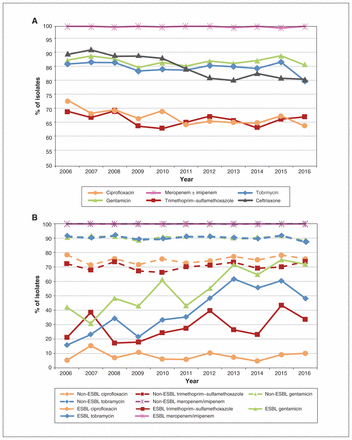
There was a significant decrease over the study period in E. coli susceptibility to ceftriaxone (p < 0.001, χ2 test for trend) and ciprofloxacin (p = 0.008, χ2 test for trend). Rates of susceptibility to other antimicrobial classes remained relatively stable (Figure 2, A). Susceptibility to ciprofloxacin, trimethoprim–sulfamethoxazole, tobramycin and gentamicin were significantly lower in ESBL-producing E. coli than in non–ESBL-producing E. coli in 2016 (Table 3). Susceptibility to ertapenem and meropenem/imipenem did not differ significantly between ESBL-producing and non–ESBL-producing E. coli. Among the former, a significant increase in aminoglycoside susceptibility was noted between 2006 and 2016 (p < 0.001 for both tobramycin and gentamicin, χ2 test for trend). Susceptibility to ciprofloxacin, trimethoprim–sulfamethoxazole, ertapenem and meropenem/imipenem remained relatively stable among ESBL-producing E. coli (Figure 2, B).
Antimicrobial susceptibilities of Escherichia coli blood culture isolates, 2006–2016. (A): Extended-spectrum β-lactamase (ESBL) producers and non–ESBL-producers combined. (B): Non–ESBL-producers and ESBL producers separately.
Susceptibility to selected antimicrobial agents of all Escherichia coli isolates, E. coli that did not produce extended-spectrum β-lactamase and E. coli that produced extended-spectrum β-lactamase, 2016
K. pneumoniae, K. oxytoca and P. mirabilis
Rates of K. pneumoniae, K. oxytoca and P. mirabilis susceptibility to ciprofloxacin, trimethoprim–sulfamethoxazole, tobramycin, gentamicin, ertapenem and meropenem/imipenem remained relatively stable over the study period. In 2016, K. pneumoniae the rates of global susceptibility to these antibiotics were 90.8%, 85.2%, 93.4%, 95.9%, 100.0% and 99.5%, respectively. For K. oxytoca, in 2016, the susceptibility rate was 100.0% for all 6 antimicrobial agents, and for P. mirabilis, the susceptibility rates were 71.4%, 77.1%, 97.1%, 97.1%, 100.0% and 100.0%, respectively. Given the small numbers of each of these species, we did not perform subset analysis of ESBL producers compared to non–ESBL-producers.
Interpretation
We found a significant increase in the proportion of ESBL-producing E. coli isolates in bloodstream infections from 2006 to 2016 in Toronto. The most striking increase was seen in the intensive care units served by our laboratory, where 25.8% of E. coli isolates were ESBL producers in 2016, compared to 11.1% in 2006. The proportion of ESBL-producing E. coli isolates also increased significantly in the emergency departments served by our laboratory, with a prevalence of 14.8% in 2016 compared to 3.3% in 2006. The fact that an increase was observed in both settings suggests that this phenomenon is not being driven by 1 ward alone and is likely being driven by outpatients as well as inpatients. In parallel with this rise in prevalence of ESBL producers, the proportion of the ST131 clone among ESBL-producing E. coli also increased, from 31.6% in 2006 to 73.0% in 2016. Interestingly, we noted a significant increase in aminoglycoside susceptibility in ESBL-producing E. coli between 2006 and 2016. This may reflect a change in circulating strains or a change in resistance-selective pressure owing to changing prescribing practices.
Clinicians should be aware of their local ESBL prevalence rates and susceptibility patterns as this informs their empiric choice of treatment, preventing delays in appropriate therapy.15,16 In 2016, 17.3% of blood culture E. coli isolates from centres served by our laboratory were ESBL-positive; this suggests that ceftriaxone may not be an appropriate empiric choice for treatment of suspected E. coli bacteremia in our patient population, especially immunocompromised patients and those presenting with sepsis, in whom agents with lower resistance rates should generally be used. Clinicians should work with their local antimicrobial stewardship teams to integrate local data, guidelines and the clinical status of patients to inform empiric treatment decisions.
Our finding of increasing ESBL prevalence is in keeping with the global trend of rising antimicrobial resistance among gram-negative bacteria.22 The importance of regional data is illustrated by the study by Peirano and colleagues,12 which showed a prevalence of ESBL production of 14% among E. coli blood culture isolates in the Calgary region in 2010, in contrast to the rate of 2.9% among E. coli isolated from all specimen types in the same year reported as part of the CANWARD study.11 There is a paucity of published data on ESBLs from the Toronto region. Lowe and colleagues23 noted an increase in the incidence rate of ESBL-positive clinical cultures from 2005 to 2009. The CANWARD study includes organisms from sentinel sites across Canada, including our laboratory, but only the first 10 blood culture isolates, (not limited to E. coli) per month in the first 10 months of the year are included, and the isolates sent from our laboratory reflect less than 10% of all blood culture isolates each year and represent less than 10% of the CANWARD data. Thus, the current study provides important data on trends in ESBL production by E. coli and other organisms in the Toronto region.
The increasing prevalence of ESBL-producing E. coli that we observed in our centre is likely related to the globally increasing prevalence of ST131 E. coli.11,12,24 This is corroborated by our finding of an increase in the frequency of the ST131 clonal type within ESBL-producing E. coli, from 31.6% in 2006 to 73.0% in 2016. ST131 E. coli is spreading worldwide, likely related to its multidrug resistance and possibly to virulence factors that provide a survival advantage.8 There is a temporal correlation of a rise in ESBL prevalence after the introduction of ST131 E. coli into a region,7 which likely contributes to regional differences in ESBL prevalence rates.
Limitations
Our data represent local ESBL prevalence rates, reflecting the patient population serviced by the centres supported by our laboratory, which include a large tertiary care transplantation centre, immunocompromised hematology cancer centre and oncology program in a large cosmopolitan city. It is likely that smaller community hospitals or centres that provide care to unique patient cohorts would show different rates. Our data are also from a predominantly adult cohort and cannot provide insight to rates among pediatric patients. Furthermore, aside from a polymerase chain reaction assay for ST131 characterization, sequencing or other molecular typing was not performed on isolates, which limits our ability to fully describe the epidemiologic features of the increase in ESBL-producing E. coli in our region. In addition, 4 isolates from 2016 were not assessed for ESBL production owing to lack of viability of the glycerol stock or presumed loss of the ESBL-resistance plasmid, and 10 ESBL-producing E. coli isolates were excluded from ST131 analysis owing to lack of viability of the glycerol stock; however, given the small numbers of isolates excluded per year, we do not believe this introduced bias into the observed trends. Finally, we cannot comment on where patients may have acquired ESBL-producing E. coli, as information on travel history and history of past hospital admissions was not assessed.
Conclusion
Our study describes recent ESBL trends in blood cultures from a Canadian region. The significant increase in the proportion of ESBL-producing E. coli in bloodstream infections in our region temporally linked with the rise in the ST131 clonal type of E. coli highlights the ongoing threat posed by antimicrobial-resistant organisms. Recognition of this significant increase is important to inform empiric antibiotic treatment of bloodstream infections. Further investigations are needed to elucidate the epidemiologic features of ESBL-producing E. coli in order to inform modification of practices aimed at reducing the prevalence and limiting the spread of ESBL-producing organisms.
Footnotes
Competing interests: Susan Poutanen has received honoraria for involvement in advisory boards and speaking events as well as travel reimbursement for speaking events from Merck Pharmaceuticals, advisory board honoraria from Paladin Labs and Verity Pharmaceuticals, grants and an advisory board honorarium from Accelerate Diagnostics, research support from Bio-Rad Laboratories, nonfinancial research support from bioMérieux and travel reimbursement from Copan Diagnostics, all outside the submitted work. No other competing interests were declared.
This article has been peer reviewed.
Contributors: Sophie Mineau and Robert Kozak contributed equally to this work. Sophie Mineau, Robert Kozak and Susan Poutanen conceived and designed the study, analyzed and interpreted the data, and drafted the manuscript. Sophie Mineau, Robert Kozak, Melissa Kissoon, Aimee Paterson, Anthony Oppedisano, Firas Douri and Kate Gogan performed the laboratory work and collected the data. Barbara Willey optimized the laboratory protocols. Allison McGeer contributed to the analysis and interpretation of the data. All of the authors critically revised the manuscript for important intellectual content, approved the final version to be published and agreed to be accountable for all aspects of the work.
Supplemental information: For reviewer comments and the original submission of this manuscript, please see www.cmajopen.ca/content/6/4/E580/suppl/DC1.
References
- Copyright 2018, Joule Inc. or its licensors