Abstract
Background: Surveillance of antimicrobial resistance is essential to mitigate its impact on population health and inform local empiric treatment practices. Our aims were to evaluate urine culture specimen susceptibility from a range of diverse settings and describe antibiotic susceptibility across all organisms and compare susceptibilities to that of Escherichia coli alone.
Methods: In this descriptive cohort study, we measured the prevalence of organisms in urine culture specimens using linked province-wide administrative databases. Using positive urine cultures collected in Ontario between Jan. 1, 2016, and Dec. 31, 2017, we measured susceptibility to 6 classes of antibiotics using a weighted antibiogram for all organisms compared with E. coli alone.
Results: We included 689 497 cultures derived from 569 399 patients and 879 778 test orders for specimens. For all organisms, the rates of susceptibility in the outpatient, inpatient and long-term care settings were 49.3%, 42.8% and 39.2%, respectively, for ampicillin; 83.1%, 72.7% and 69.7%, respectively, for nitrofurantoin; 80.3%, 64.8% and 73.1%, respectively, for trimethoprim–sulfamethoxazole; 87.2%, 74.1% and 66.2%, respectively, for ciprofloxacin; 90.6%, 73.6% and 85.1%, respectively, for aminoglycosides; and 82.6%, 57.5% and 73.5%, respectively, for cefazolin. We found resistance to 3 or more antibiotic classes in 20.6% of episodes for all organisms compared with 14.0% for E. coli alone. The average absolute difference in antibiotic susceptibility between all organisms and E. coli across all drugs was lowest in the outpatient setting (6.2%) and highest in the inpatient setting (14.6%).
Interpretation: In this study, urinary organism prevalence and antimicrobial susceptibility varied across health care settings and patient populations, with implications for both antimicrobial resistance surveillance and clinical decision-making. Weighted antibiograms may be most useful for guiding empiric treatment of urinary infections in inpatient settings where the diversity of infectious organisms is higher than in the community.
Antimicrobial resistance is a worsening threat to both individual and population health worldwide. Increasing levels of antimicrobial resistance threaten to limit available options for treating serious infections, as well as making infections that were once considered minor or a nuisance more problematic. At the population level, increased emergence and transmission of antimicrobial resistance can lead to strain on health care systems, and substantial social and economic impacts.1,2
Large electronic repositories allow for more comprehensive data to be collected and analyzed for a wider range of organisms across health care settings. The simplest approach is to use an antibiogram. Antibiograms are tables that provide the susceptibility for a variety of organism–drug combinations to support optimal empiric therapy, typically in a specific setting (e.g., hospital wide or intensive care unit).3
An improvement on the traditional antibiogram is the weighted-incidence syndromic combination antibiogram (WISCA), which is particularly well suited for data from a single body site and has been described previously.4–6 This method leverages the relative contributions of different organisms that cause a clinical syndrome such as a urinary tract infection (UTI) at the population level, and pools organism- and drug-specific susceptibilities to provide a likelihood of appropriate antibiotic coverage before the organism is identified in culture.4 A WISCA simplifies a traditional antibiogram by providing a single susceptibility estimate for a given drug because the susceptibility for each organism is pooled and weighted based on their incidence, making it easier to select a more appropriate empiric antibiotic before the availability of patient-specific culture results.4,5 The use of WISCAs in clinical practice is associated with more appropriate antibiotic empiric selection,5,7,8 but this method may also be useful for surveillance of antimicrobial resistance in UTIs at the population level and informing future guideline development and interventions.
In this study, we aimed to measure the prevalence of antibiotic susceptibility in urinary isolates in Ontario. We compared overall susceptibility obtained from an advanced WISCA approach (i.e., an all-organism weighted approach) to results generated for Escherichia coli alone by patient setting for commonly reported antibiotics.
Methods
Study design and setting
We conducted this descriptive cohort study in Ontario, Canada’s most populous province, using data from a population-based electronic repository of laboratory results consisting of 121 reporting sites across 14 health regions serving a population of 14 million.9,10 The reporting sites included hospital, community and public health laboratories.
We reported the study using the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) reporting guideline checklist.11
Population
We measured the prevalence of urinary organisms and susceptibility from testing episodes using currently available data (from Jan. 1, 2016, to Dec. 31, 2017). All patients in Ontario regardless of age or health care setting were eligible for inclusion.
Data sources
We obtained study data from linked population-wide administrative data sets housed at ICES. ICES is an independent, nonprofit research institute whose legal status under Ontario’s health information privacy law allows it to collect and analyze health care and demographic data, without consent, for health system evaluation and improvement.
Urine culture and susceptibility data were captured from the Ontario Laboratories Information System (OLIS), a large centralized electronic repository for laboratory results for public health, hospital and outpatient facilities; more than 95% of clinical microbiology laboratories in the province report culture and susceptibility results into OLIS.12,13 The OLIS repository was introduced in Ontario in 2007 to allow timely access to laboratory results for clinical purposes. Laboratory participation gradually increased over the following decade to provide wide coverage across the province. Data extracted from the repository included test request (ordering), specimen (source), results and organism data.
We obtained demographics data for the study population from the Registered Persons Database (RPDB), Canadian Institute for Health Information Discharge Abstract Database (CIHI-DAD) and Continuing Care Reporting System — Long-Term Care (CCRS-LTC). These data sets are widely validated and were linked using unique encoded identifiers and analyzed at ICES14–16 (Appendix 1, Supplementary Table 1, available at www.cmajopen.ca/content/10/4/E1044/suppl/DC1).
Urine culture antibiotic susceptibility, by organism and patient characteristics
We included patient age, sex, health region, health care setting and year at the time of specimen collection in addition to the organism isolated. We categorized health care settings as inpatient (acute care admission in hospital), long-term care and outpatient (community physicians or emergency department visits) settings.
Outcomes
Antibiotic susceptibility interpretations for select drugs (ampicillin, nitrofurantoin, trimethoprim–sulfamethoxazole, ciprofloxacin, cefazolin, gentamicin and tobramycin) as reported by the laboratory to clinicians were the primary outcomes in our study. Because of the variability in reporting practices of the included laboratories, we used a stepwise process to impute missing outcomes and retain the maximum number of records.
Raw antibiotic susceptibility
We retrieved antibiotic susceptibility interpretations for the drugs listed above from OLIS for records with a valid culture result. We grouped intermediate resistance and resistant results into a nonsusceptible result category.6 We coded missing susceptibility results as such and retained them. We combined results for gentamicin and tobramycin into the aggregate class of aminoglycosides, whereby resistance to either drug indicated resistance to the class, given the similarity of these 2 agents in terms of their spectrum of activity.
Direct rule-based imputation
To account for the variability in laboratory reporting of antibiotic susceptibility results, we developed direct imputation rules for deriving missing susceptibility results that are known from organisms’ microbiological characteristics, such as intrinsic susceptibility (always susceptible regardless of presence or absence of reported results), intrinsic resistance (always resistant regardless of presence or absence of reported results) and cross-resistance between antibiotic classes (if the organism is susceptible or resistant to 1 agent, it will always be susceptible or resistant to another specific agent; Appendix 1, Supplementary Table 2). For example, certain laboratories did not report ampicillin susceptibility results for Klebsiella pneumoniae because it is known to be intrinsically resistant. These rules were based on standard bacteriological testing and reporting recommendations that help to infer susceptibility or resistance across more than 1 agent based on common resistance mechanisms.6,17
Setting-specific organism distribution for urine culture episodes in Ontario
Model-based imputation
We used 3 logistic regression models for imputing missing susceptibility results from organism–drug combinations that could not be determined with direct rule-based imputation (Appendix 1, Supplementary Table 3). The first model included all patient characteristics (age, sex, setting, health region and organism) and susceptibility results for the 5 drugs that were not being imputed. We performed the logistic regression on the subset of records for which the organism–drug susceptibility result of interest was not missing, and the modelled log-odds were converted to a probability of a susceptible result. In organism–drug combinations where this model did not converge, we attempted a second regression model on the same subset using only the patient characteristics and no drugs. We applied an intercept-only logistic model to the remaining organism–drug combinations that did not converge owing to low numbers or nonvariable outcomes. The resulting modelled probabilities were then applied to the subset for which the organism–drug susceptibility result was missing, and a result randomly assigned to each record according to the modelled probability.
Statistical analysis
For each antibiotic studied, we calculated overall percent susceptibility by organism, age group, sex, health care setting, year and health region. We also determined the distributions for health care setting–specific organism infections for all urine culture episodes. We determined the weighted susceptibility of all organisms in combination. We also calculated differences between percent susceptibility between susceptibility for all organisms combined and E. coli alone, by drug and patient setting. We used SAS version 9.4 to perform all analyses.
Ethics approval
The use of the data in this project is authorized under section 45 of Ontario’s Personal Health Information Protection Act and does not require review by a research ethics board.
Results
Our study included results for urine culture antibiotic susceptibility from 689 497 unique annual patient/organism testing episodes derived from 569 399 patients and 879 778 test orders for specimens collected between Jan. 1, 2016, and Dec. 31, 2017 (Figure 1). Of the positive results for urine culture, we included only monomicrobial cultures, and 57.9% of those were excluded because no valid susceptibility testing was available. Of the remaining orders with susceptibility testing, cefazolin was the most common missing antibiotic (10.8%) and nitrofurantoin was the most complete, with results missing in only 2.5% of cultures. In all, 79% of urine cultures with valid susceptibility testing available had complete susceptibility results for all tested agents before imputation. Rule-based imputation added results for 9.7% of testing episodes, and model-based imputation added results for the remaining 11.3% of orders. We excluded records if they could not be linked to a unique encoded patient identifier (0.2%). When multiple records existed for a patient with the same organism and year (21.6%), we aggregated results to 1 unique episode by retaining the first instance.
Flow chart showing the inclusion selection process for urine culture episodes.
Most culture episodes (85.6%) were obtained from female patients and were primarily from the 18–64 years age group (50.7%) (Table 1). Patients aged less than 18 years and those 65 years or older represented 5.6% and 43.7% of the study population, respectively. We found that most episodes were associated with outpatient settings (88.4%), followed by acute care (5.9%) and long-term care (5.7%). The proportion of episodes contributed by each of the 14 health regions ranged from 1.0% to 14.0%.
Escherichia coli was the most commonly isolated organism (found in 72.2% of patient testing episodes), followed by K. pneumoniae (8.9%), Proteus mirabilis (4.0%), Pseudomonas aeruginosa (1.6%) and Citrobacter koseri (1.5%). Each of the remaining 18 species or species groups individually accounted for less than 1.5% of all episodes. The proportion of episodes attributable to E. coli varied by setting, with 49.0% in inpatient, 59.1% in long-term care and 74.6% in outpatients (Table 2).
We found that antibiotic susceptibility was highest in the outpatient setting for all drugs, and the difference between the highest and lowest susceptibility by setting ranged from 10.1% for ampicillin (49.3% outpatient v. 39.2% long-term care) to 25.1% for cefazolin (82.6% outpatient v. 57.5% inpatient). Susceptibility was lowest in the long-term care setting for ampicillin, nitrofurantoin and ciprofloxacin, and lowest in the inpatient setting for trimethoprim–sulfamethoxazole, aminoglycosides and cefazolin.
Susceptibility was highest in female patients for all drugs, with the greatest difference between males and females found for cefazolin (83.7% v. 62.4%). Patients aged 65 years or older had the lowest susceptibility across all drugs except for ciprofloxacin: patients aged less than 18 years were 72.9% susceptible compared with 81.8% susceptible among those aged 65 years or older. Ciprofloxacin showed the biggest spread in susceptibility between health regions at 8.5% (88.0% in region 2 v. 79.5% in region 5), whereas nitrofurantoin varied by only 3.4% between the most (83.9%, region 3) and least (80.5%, region 5) susceptible regions.
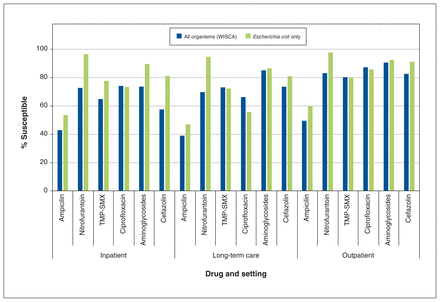
We found that organisms were most susceptible to aminoglycosides (89.3%) and least susceptible to ampicillin (48.3%), and there was resistance to 3 or more drugs for 20.6% of patient testing episodes. The difference between antibiotic susceptibility for all organisms combined versus only E. coli varied by both drug and setting (Figure 2). Antibiotic susceptibility for all organisms combined (i.e., allorganism susceptibility) was lower than E. coli alone across all settings for ampicillin, nitrofurantoin, aminoglycosides and cefazolin. The difference was much greater in inpatient specimens compared with long-term care and outpatient specimens for trimethoprim–sulfamethoxazole (−12.9% inpatient, 0.8% long-term care and 0.5% outpatient), aminoglycosides (−16.0% inpatient, −1.4% long-term care and −1.9% outpatient) and cefazolin (−23.7% inpatient, −7.4% long-term care and −8.6% outpatient). Only ciprofloxacin had higher antibiotic susceptibility for all organisms versus E. coli across all settings, with the difference being greater in the long-term care setting (10.6%) compared with inpatient (0.8%) and outpatient (1.5%) specimens. Combined susceptibility for nitrofurantoin was much lower than E. coli alone across all settings with differences of −23.8%, −25.0% and −14.5% for inpatient, long-term care and outpatient settings, respectively. The average absolute difference between all organisms and E. coli across all drugs was lowest in the outpatient setting (6.2%) and highest in the inpatient setting (14.6%).
Urine culture antibiotic susceptibility by setting and drug for all organisms weighted-incidence syndromic combination antibiogram (WISCA) versus Escherichia coli only. Note: TMP–SMX = trimethoprim–sulfamethoxazole.
Interpretation
We conducted a comprehensive, population-based antibiotic susceptibility profile for urine culture isolates across community (i.e., outpatient), long-term care and acute care (i.e., inpatient) settings. The inclusion of results from these diverse patient settings allowed us to evaluate where allorganism derived susceptibilities differed compared with E. coli susceptibilities alone. It is important to note that, in the outpatient setting, 80%–85% of positive urine cultures were susceptible to nitrofurantoin, trimethoprim–sulfamethoxazole, cephalexin and ciprofloxacin.
We also found that the proportions susceptible were most similar across all drugs between the 2 methods in the outpatient setting, which was largely driven by the higher proportion of episodes caused by E. coli. This is in line with previous studies that described E. coli as being more common in outpatient than in inpatient UTIs.18–21 Although E. coli is commonly tracked as a marker for urinary gram-negative susceptibility in outpatients,22,23 the prevalence of species other than E. coli may contribute substantially to overall resistance. Although other uropathogens are less common, they may have higher rates of antimicrobial resistance compared with E. coli.
All-organism derived antibiograms may provide a more accurate assessment of antibiotic susceptibility, particularly in health care facilities; however, antibiograms may be less useful in the outpatient setting where infections caused by E. coli predominate. For example, 83.1% of isolates were susceptible to nitrofurantoin (a first-line antibiotic for uncomplicated cystitis)24 using the all-organism derived antibiogram compared with only 69.7% and 72.7% of isolates in long-term care and hospitals, respectively, which suggests a more tailored approach to selection of empiric treatment may be needed in these settings. The 2011 iteration of the Infectious Diseases Society of America and the European Society for Microbiology and Infectious Diseases guideline for uncomplicated UTIs recognized the importance of accounting for local resistance rates for all potential pathogens when selecting empiric antibiotic therapy.24 For example, the empiric uses of trimethoprim–sulfamethoxazole and fluoroquinolones are based on local uropathogen resistance thresholds of no more than 20% and 10%, respectively, rather than the most common pathogen (E. coli). These recommendations support the combined all-organism approach to estimate antimicrobial susceptibility across all potential pathogens in a given population.
In addition to health care settings, we found differences in urinary antibiotic susceptibility across age groups (more resistance in children and adults aged 65 yr and older), sex (more resistance in male patients) and region. Higher antibiotic use at the extremes of age,25 as well as increasing prevalence pathogens other than E. coli among older age groups are likely to drive age-specific differences and patterns of antibiotic susceptibility. The higher prevalence of antimicrobial resistance in males than in females may be reflected by the more complicated nature of UTIs in males (UTI is more likely to be caused by a functional or anatomic abnormality),26 as well as the predominance of younger females who were outpatients in our cohort.
Regional variation in demographics and, more importantly, antibiotic prescribing practices25 may be key factors in explaining regional differences in antibiotic resistance among urinary pathogens. An interactive urinary antibiogram created using 2016–2017 data is available from Public Health Ontario.27 The antibiogram includes detailed organism–drug susceptibility results by region, care setting and age group. Given the heterogeneity in antibiotic resistance, harnessing population-level susceptibility data and tailoring all-organism derived antibiograms to the local population may be a helpful approach to support improved antibiotic decision-making. The comprehensiveness of these data could facilitate the development of centralized tools that would allow practitioners to tailor empiric therapy based on patient characteristics predictive of resistance.
Strengths of our study include the comprehensive population-wide data set that included 689 497 urine culture episodes, and imputation and modelling strategy. This approach to filling in missing values relied on regression modelling in contrast to expert opinion and literature review as used in other WISCAs.4 Regression modelling provides an advantage by imputing susceptibility results based on the distribution of nonmissing results specific to our cohort as opposed to extrapolating from studies of potentially noncomparable populations.
Limitations
A key limitation to the use of centralized laboratory data from numerous laboratories is that only 42.1% of all positive urine culture orders were associated with any susceptibility results for the selected antibiotics. Potential causes include laboratory policies regarding susceptibility testing on repeat specimens, laboratories reporting susceptibility results on nonstandard antibiotics, duplicate culture orders introduced by reference testing, missing susceptibility results from polymicrobial cultures and coding errors. As the OLIS repository comprises almost all results reported back to clinicians in the province, this characteristic should not affect the representativeness of the data we presented. Although the populations differ slightly, the estimated susceptibility results from our study (e.g., E. coli susceptibility to nitrofurantoin is 98%) align with those found in other sources such as the Ontario Hospital Antibiogram27,28 (E. coli susceptibility to nitrofurantoin is 97%), which reports data provided directly from hospital antibiograms, and LifeLabs29 (E. coli susceptibility to nitrofurantoin is 97% in the Toronto region), which reports data from a large private laboratory in Canada.
A large proportion of the culture episodes are likely to have come from asymptomatic bacteriuria, which generally does not require treatment. Therefore, a limitation to these administrative data is the inability to ascertain symptoms and confirm a diagnosis of UTI in these patients. Currently, these data include the years 2016 and 2017 and susceptibilities may have changed in subsequent years. Despite these limitations, we were able to report antibiotic susceptibility on a substantial number of urine culture episodes with the ability to compare susceptibility across several patient settings for a variety of predominant organisms.
Conclusion
Prevalence of urinary organisms and antimicrobial susceptibility varies across health care settings and patient populations. This variability has implications for both surveillance of antimicrobial resistance and clinical decision-making. Given the difference in prevalence of urinary E. coli between settings, use of an all-organism derived antibiogram may more accurately reflect resistance compared with E. coli alone, particularly in long-term care and inpatient settings. Future efforts are needed to evaluate the impact of this approach on the appropriateness of empiric therapy and patient outcomes.
Footnotes
Competing interests: None declared.
This article has been peer reviewed.
Contributors: Alex Marchand-Austin collected and analyzed the data, interpreted the results and drafted the article. Samantha Lee contributed to the collection and interpretation of the data, as well as drafted and critically revised the article. Bradley Langford, Nick Daneman and Kevin Brown conceptualized and designed the study, and collaborated and contributed to the analysis and interpretation of the data, and critically revised the article. Derek MacFadden, Christina Diong, Kevin Schwartz, Vanessa Allen, Jennie Johnstone, Samir Patel and Gary Garber contributed to the conception of the study, interpreted the data and critically revised the article for important intellectual content. All of the authors gave final approval of the version to be published and agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Funding: This work was supported by the Canadian Institutes of Health Research (funding reference no. 159503).
Data sharing: The data set from this study is held securely in coded form at ICES. Although legal data-sharing agreements between ICES and data providers (e.g., health care organizations and government) prohibit ICES from making the data set publicly available, access may be granted to those who meet prespecified criteria for confidential access, available at https://www.ices.on.ca/DAS (email: das{at}ices.on.ca). The full data set creation plan and underlying analytic code are available from the authors upon request, with the understanding that the computer programs may rely upon coding templates or macros that are unique to ICES and are therefore either inaccessible or may require modification.
Disclaimer: This study was supported by ICES, which is funded by an annual grant from the Ontario Ministry of Health and the Ministry of Long-Term Care. Parts of this material are based on data or information compiled and provided by the Canadian Institute for Health Information (CIHI). However, the analyses, conclusions, opinions and statements expressed in the material are those of the authors, and not necessarily those of CIHI.
Supplemental information: For reviewer comments and the original submission of this manuscript, please see www.cmajopen.ca/content/10/4/E1044/suppl/DC1.
This is an Open Access article distributed in accordance with the terms of the Creative Commons Attribution (CC BY-NC-ND 4.0) licence, which permits use, distribution and reproduction in any medium, provided that the original publication is properly cited, the use is noncommercial (i.e., research or educational use), and no modifications or adaptations are made. See: https://creativecommons.org/licenses/by-nc-nd/4.0/
References
- © 2022 CMA Impact Inc. or its licensors