Abstract
Background: Surveillance of antimicrobial resistance is vital to guiding empirical treatment of infections. Collating and reporting routine data on clinical isolate testing may offer more timely information about resistance patterns than traditional surveillance network methods.
Methods: Using routine microbiology testing data collected from the Bacteremia Antibiotic Length Actually Needed for Clinical Effectiveness retrospective cohort study, we conducted a descriptive secondary analysis among critically ill patients in whom bloodstream infections had been diagnosed in 14 intensive care units (ICUs) in Canada. The participating sites were located within tertiary care teaching hospitals and represented 6 provinces and 10 cities. More than 80% of the study population was accrued from 2011-2013. We assessed the epidemiologic features of the infections and corresponding antimicrobial susceptibility profiles. Susceptibility testing was done according to Clinical Laboratory Standards Institute guidelines at accredited laboratories.
Results: A total of 1416 pathogens were isolated from 1202 patients. The most common organisms were Escherichia coli (217 isolates [15.3%]), Staphylococcus aureus (175 [12.4%]), coagulase-negative staphylococci (117 [8.3%]), Klebsiella pneumoniae (86 [6.1%]) and Streptococcus pneumoniae (85 [6.0%]). The contribution of individual pathogens varied by site. For 13 ICUs, gram-negative susceptibility rates were high for carbapenems (95.4%), tobramycin (91.2%) and piperacillin-tazobactam (90.0%); however, the proportion of specimens susceptible to these agents ranged from 75.0%-100%, 66.7%-100% and 75.0%-100%, respectively, across sites. Fewer gram-negative bacteria were susceptible to fluoroquinolones (84.5% [range 64.1%-97.2%]). A total of 145 patients (12.1%) had infections caused by highly resistant microorganisms, with significant intersite variation (range 2.6%-24.0%, χ2 = 57.50, p < 0.001).
Interpretation: We assessed the epidemiologic features of bloodstream infections in a geographically diverse cohort of critically ill Canadian patients using routine pathogen and susceptibility data extracted from readily available microbiology testing databases. Expanding data sharing across more ICUs, with serial measurement and prompt reporting, could provide much-needed guidance for empiric treatment for patients as well as system-wide prevention methods to limit antimicrobial resistance.
Bloodstream infections are associated with considerable morbidity, mortality and cost. In North America, it is estimated that 575 000-677 000 episodes and 79 000-94 000 deaths per year are attributable to nosocomial bloodstream infections; these infections are the sixth- and seventh-leading causes of death in Canada and the United States, respectively.1 Current, accurate data on the epidemiologic features of bloodstream infections and antimicrobial susceptibility profiles are needed to guide appropriate empirical antimicrobial treatment and improve patient outcomes.2 Data on the epidemiologic characteristics of bloodstream infections are especially vital in intensive care unit (ICU) settings, where these infections are most common, antibiotic resistance is most prevalent, and timely, appropriate antibiotic therapy is necessary to prevent complications and death.3,4
Epidemiologic data on bloodstream infections are usually provided by national microbiology surveillance networks. The Canadian Ward Surveillance Study5 is a typical example of such a network. Clinical isolates are collected from infected inpatients and outpatients at a number of tertiary care medical centres across Canada. Each centre is asked to provide a minimum number of respiratory, urine, wound and blood isolates per month based on set criteria as well as limited patient demographic data (age, sex, hospital location and specimen source). The isolates are sent to a central coordinating laboratory for antimicrobial susceptibility testing, and results are analyzed centrally to assess the national epidemiologic features of infections, including resistance patterns. These monitoring networks offer broad geographic coverage, standardized testing methods and measurement of exact minimum inhibitory concentrations; however, these strengths often come at the expense of timeliness, as they rely on referral of microbial isolates to a centralized laboratory, direct microbiology laboratory testing and reporting into a centralized database.6 Thus there is a scarcity of current data on the profile of pathogens and the antimicrobial susceptibility profiles of pathogens causing serious bacterial infections in Canada.
Routine diagnostic data from local hospital laboratories, in contrast, may offer a timely and direct source of data on epidemiologic features and susceptibility profiles because they can be extracted from available data.6 Although there may be administrative delays in establishing a network based on routinely available data, once established, such a network would be able to more quickly generate serial updates by circumventing the need to gather, transport and process biologic specimens. These routine databases also offer the added benefit of being potentially linkable to detailed clinical and outcome data, which are more difficult to capture in surveillance networks, and, in addition, provide the opportunity to assess trends in epidemiologic and susceptibility data at a local level, which can help inform decision-making at hospitals that contribute data. The European Antimicrobial Resistance Surveillance Network has harnessed routine culture data to assess periodic trends in antimicrobial-resistant organisms across Europe.7
The proof-of-concept objective of our study was to use routine microbiology testing data to describe the epidemiologic features of bloodstream infections and corresponding antimicrobial susceptibility profiles from a recent Canadian multicentre cohort of critically ill patients.
Methods
Study sites and patients
This was a secondary analysis of the Bacteremia Antibiotic Length Actually Needed for Clinical Effectiveness study, a multisite retrospective cohort study of critically ill patients in whom bloodstream infection had been diagnosed in 14 ICUs across Canada.8 The participating sites were located within tertiary care teaching hospitals and represented 6 provinces and 10 cities (Appendix 1, available at www.cmajopen.ca/content/4/4/E569/suppl/DC1). Patients who had a blood culture that grew a pathogenic organism during their ICU stay were eligible for inclusion in the study. The cohort was accrued by reviewing patient records prior to December 2013 to identify the most recent consecutive critically ill patients who had bloodstream infections, up to a maximum of 100 patients per site. Seven sites contributed data for 100 patients, 5 sites contributed data for 82 to 99 patients, 1 site contributed data for 38 patients, and 1 site contributed data for 5 patients. More than 80% of the study population was accrued from 2011-2013. Only 1 episode of bacteremia was included per patient, but all organisms isolated in blood culture sets over the first 24-hour period from the index blood culture were considered to be contributors to the index bacteremia.
Because the cohort was designed to examine prevailing treatment durations for bacteremia without a deep-seated focus, patients were excluded if they had endocarditis, osteomyelitis, septic arthritis, an undrained abscess or unremoved prosthetic material. Patients were also excluded if they had a single culture containing only a common contaminant (coagulase-negative staphylococci, Corynebacterium spp, Bacillus spp, Propionobacterium spp, Aerococcus spp or Micrococcus spp).9-12
Data collection and measures
Experienced research coordinators affiliated with the Canadian Critical Care Trials Group at each ICU retrospectively abstracted data on cultured pathogen(s) and susceptibility testing results, source of bloodstream infection, antimicrobial treatment and clinical outcomes. The data were entered into a secure Web-based electronic case report form, and checks were made to minimize missing and invalid data.
Susceptibility testing was done according to Clinical Laboratory Standards Institute guidelines at accredited laboratories.13 Infections were classified as community-acquired if they had been diagnosed by means of a blood culture obtained within 48 hours of hospital admission, hospital-acquired if obtained more than 48 hours after hospital admission, and ICU-acquired if obtained more than 48 hours after admission to the ICU. The source of infection was based on a review of the history, physical and laboratory findings, and clinician notes. The patient's Acute Physiology and Chronic Health Evaluation II score was noted; this is a measure of severity of disease (range 0-71) calculated within 24 hours of ICU admission that is based on initial values of 12 routine physiologic measurements, age and previous health status, with higher scores indicating more severe disease.14 Last, we defined highly resistant microorganisms using a modified version of the definition proposed by de Smet and colleagues:15 methicillin-resistant Staphylococcus aureus; vancomycin-resistant Enterococcus spp; penicillin-resistant Streptococcus pneumoniae; extended-spectrum β-lactamase-producing Enterobacteriaceae; carbapenem-resistant Enterobacteriaceae; carbapenem-resistant Acinetobacter spp; Enterobacteriaceae resistant to at least 2 of fluoroquinolones, aminoglycosides or trimethoprim-sulfamethoxazole; Acinetobacter spp resistant to at least 2 of fluoroquinolones, aminoglycosides or ceftazidime; or non-Enterobacteriaceae resistant to at least 3 of fluoroquinolones, aminoglycosides, carbapenems, ceftazidime or piperacillin.
Statistical analysis
We conducted all analyses at the pathogen level (pathogen description, susceptibility profiles) with the exception of describing the patients and the prevalence of highly resistant microorganisms in this population, which was done at the patient level. To examine variability in the proportion of patients with specific pathogens and antibiotic resistance profiles across ICUs, we used the Pearson χ2 test or Fisher exact test for comparisons and excluded the site that contributed data for only 5 patients to the cohort. All other analyses included all patients. We used binomial exact methods to calculate 95% confidence intervals. For the antimicrobial susceptibility analysis, we defined isolates as either susceptible or resistant to an antimicrobial; isolates of intermediate susceptibility were categorized as resistant. We conducted analyses using Stata v12.
Ethics approval
Approval was granted by the research ethics boards of all participating ICUs, and informed consent was waived.
Results
Patient description
A total of 1202 patients were included in the analysis. They were an average of 60 (standard deviation 17) years old (range 16-95 years), were mostly male (748 [62.2%]) and had been admitted for medical (933 [77.6%]), surgical (136 [11.3%]) or trauma-related (72 [6.0%]) reasons. The patients had a mean Acute Physiology and Chronic Health Evaluation II score of 23 (standard deviation 9) at admission, and most (1110 [92.3%]) had at least 1 comorbid condition. Of the 1202, 479 (39.8%) died. Half (602 [50.1%]) of the infections were community-acquired, 220 (18.3%) were hospital-acquired, and 380 (31.6%) were ICU-acquired. Pneumonia/lung infections (453 patients [37.7%]), urinary tract infections (243 [20.2%]) and vascular catheters (241 [20.0%]) were the most common sources of the infections. For the remaining patients, the source of bacteremia was intra-abdominal (189 [15.7%]), hepatobiliary (78 [6.5%]), skin or soft tissue (97 [8.1%]), other infections (62 [5.2%]) or unknown (186 [15.5%]). Most patients (1025 [85.3%]) were infected with 1 unique organism, but 177 patients (14.7%) had polymicrobial infections. Mortality rates varied significantly by acquisition setting, with the highest rates among patients with hospital-acquired (52.9%) and ICU-acquired (44.8%) infections and the lowest rate among those with community-acquired infections (32.7%) (p < 0.001). Mortality rates were similar for patients with monomicrobial infection and those with polymicrobial infection (39.6% v. 41.2%) (p = 0.682).
Pathogen description
In total, 1416 pathogens were isolated from the 1202 patients: 558 (39.4%) were gram-negative bacilli, 667 (47.1%) were gram-positive cocci, 81 (5.7%) were other bacteria, and 94 (6.6%) were yeast; 16 pathogens did not have names listed in the database and so their pathogen group could not be assigned. The most frequently isolated individual pathogens, grouped by species, are shown in Table 1; the 5 most common organisms were Escherichia coli (217 [15.3%]), Staph. aureus (175 [12.4%]), coagulase-negative staphylococci (117 [8.3%]), Klebsiella pneumoniae (86 [6.1%]) and Strep. pneumoniae (85 [6.0%]). Mortality rates varied by infective organism group, from 31.5% (Klebsiella spp) to 64.5% (Candida spp).
E. coli was the most frequently isolated gram-negative bacillus for both community- and hospital-acquired infections, whereas Pseudomonas aeruginosa was more common in ICU-acquired infections (Table 1). Enterococcus spp were the most frequently isolated gram-positive organisms in ICU-acquired infections. Significantly lower proportions of coagulase-negative staphylococci were detected in patients with community-acquired (6.2%) and hospital-acquired (6.0%) infections than in those with ICU-acquired infection (13.0%) (χ2 = 18.89, p < 0.001). Similar findings were observed for Candida spp (2.9% and 8.3% v. 11.6%) (χ2 = 34.48, p < 0.001). In contrast, Strep. pneumoniae was cultured significantly more frequently among patients with community-acquired infections (11.4%) than among those with infections acquired in the hospital (1.1%) or the ICU (0.2%) (χ2 = 73.52, p < 0.001).
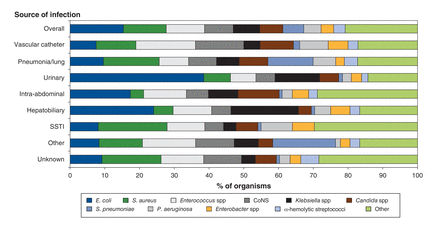
In patients in whom the source of the infection was identified to be urinary tract- or hepatobiliary-related, infections were caused by gram-negative bacteria, primarily E. coli and K. pneumoniae (Figure 1). Excluding Staph. aureus, gram-positive cocci contributed to a greater relative proportion of infections than did other organisms for other sites including vascular catheter and intra-abdominal (Figure 1).
Distribution of organisms by source of infection in critically ill patients with bloodstream infections (n = 1416). CoNS = coagulase-negative staphylococci, SSTI = skin and/or soft-tissue infection.
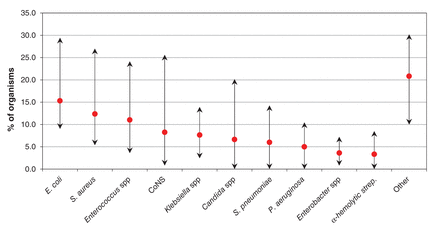
There was substantial variation in the pathogens isolated across the ICUs (χ2 = 450.66, p < 0.001) (Figure 2). The proportion of Staph. aureus isolates ranged from 5.3%-27.0%, coagulase-negative staphylococci 0.8%-25.6%, other gram-positive cocci 15.4%-33.1%, gram-negative bacilli 16.7%-50.9%, Candida spp 0%-20.2%, and other bacteria 0%-9.8%.
Variation in isolated pathogens across 13 intensive care units. Arrows represent the range in the proportion of the particular pathogen across sites, and the red dots represent the overall proportion of the pathogen isolated for all intensive care units combined. CoNS = coagulase-negative staphylococci.
Susceptibility profiles
The antimicrobial susceptibility profiles of the pathogens are shown in Table 2. Of the specimens that tested positive for Staph. aureus, 76.4% were susceptible to methicillin and 73.4% were susceptible to fluoroquinolones, but 95.0% were susceptible to trimethoprim-sulfamethoxazole, and 93.9% were susceptible to doxycycline. One-third (32.9%) of specimens positive for coagulase-negative staphylococci were susceptible to methicillin, whereas most (87.5%) were susceptible to doxycycline. Enterobacteriaceae susceptibility rates were high for aminoglycosides (90.9%-97.8%) and carbapenems (98.6%). Fewer non-Enterobacteriaceae specimens were susceptible to carbapenems (80.9%).
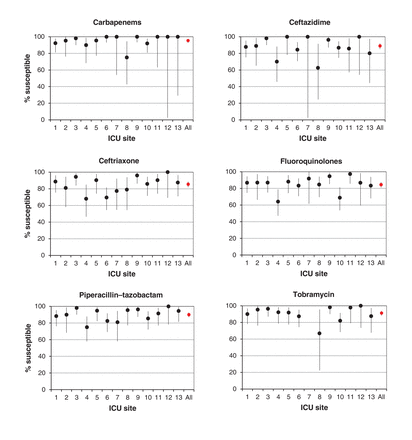
We observed variability in susceptibility profiles across the ICUs when we examined all gram-negative bacteria (Figure 3). Overall, susceptibility rates were high for carbapenems (95.4%), tobramycin (91.2%) and piperacillin-tazobactam (90.0%); however, the proportion of specimens susceptible to these agents ranged from 75.0% to 100%, 66.7% to 100% and 75.0% to 100%, respectively, across sites. Fewer gram-negative bacteria were susceptible to fluoroquinolones (overall susceptibility 84.5% [range across sites 64.1%-97.2%]).
Variation in susceptibility to antibacterial agents in patients with gram-negative bloodstream infections (n = 558) across 13 intensive care units. Error bars represent 95% confidence intervals.
Highly resistant microorganisms
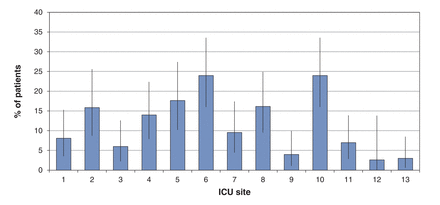
Overall, 145 patients (12.1%) had infections caused by highly resistant microorganisms, most commonly extended-spectrum β-lactamase-producing Enterobacteriaceae (54 [4.5%]) and methicillin-resistant Staph. aureus (41 [3.4%]). Thirty patients (2.5%) were infected with Enterobacteriaceae resistant to at least 2 of fluoroquinolones, aminoglycosides or trimethoprim-sulfamethoxazole, and 23 patients (1.9%) were infected with vancomycin-resistant Enterococcus spp. Four patients (0.3%) had carbapenem-resistant Enterobacteriaceae. There was significant variability across ICUs in the proportion of patients infected with highly resistant microorganisms, ranging from 2.6% to 24.0% (χ2 = 57.50, p < 0.001) (Figure 4).
Variation in the proportion of patients with highly resistant microorganisms (n = 145) across 13 intensive care units (ICUs). Error bars represent 95% confidence intervals.
Interpretation
We assessed the epidemiologic features of bloodstream infections in a geographically diverse cohort of critically ill Canadian patients using routine pathogen and susceptibility data extracted from readily available microbiology testing databases. More than 100 unique organisms were isolated, half of which were gram-positive cocci, with significant variation in isolated organisms across acquisition settings, source of infection and ICU site. We noted high susceptibility rates (> 95%) for carbapenems against Enterobacteriaceae and for amikacin against non-Enterobacteriaceae; despite this, 1 in 8 patients had infections caused by highly resistant microorganisms, with the proportion varying significantly across sites.
The 5 most frequently isolated pathogens in our study, E. coli, Staph. aureus, coagulase-negative staphylococci, K. pneumoniae and Strep. pneumoniae, were among the 10 most frequently isolated pathogens in blood cultures between 2007 and 2009 in the Canadian Ward Surveillance Study.16 However, infection due to Candida spp was relatively more frequent in our cohort than in the Canadian Ward Surveillance Study. This difference likely reflects the different settings for data collection, as our study focused exclusively on critical care settings, where Candida spp infections are more prevalent.17 Similarly, our antimicrobial susceptibility findings were consistent with those of other Canadian studies16 and with American and European critical care data from the SENTRY Antimicrobial Surveillance Program.3 We did, however, observe higher proportions of methicillin-resistant Staph. aureus and vancomycin-resistant enterococci infections in our cohort compared to the Canadian Nosocomial Infection Surveillance Program.18-20 This may reflect the inclusion of more recent data in our study, as colonization and infection rates of vancomycin-resistant enterococci are increasing in Canada over time.19 The differences may also be attributable to differing patient populations, as resistance rates are higher in critical care settings.21 Despite this, we identified only 4 patients with carbapenem-resistant Enterobacteriaceae, from 4 different ICU sites, which suggests sporadic and less frequent occurrence than in Europe.22 These differences may be due in part to increased detection and reporting in Europe:22 a recent study showed that 14% of infections due to carbapenem-resistant Enterobacteriaceae may go undetected in Ontario based on current Clinical Laboratory Standards Institute recommendations.23
We found significant variability in the isolated causative organisms for patients' bloodstream infections, susceptibility profiles and proportion of highly resistant microorganisms across ICU sites. Similar heterogeneity has been reported recently between hospitals.4,24 Hypothesized hospital-level factors contributing to variation in highly resistant microorganisms include differing infection prevention and control practices, antimicrobial use practices, hospital size, staffing (nurse ratios, infection prevention and/or hospital epidemiology staffing) and characteristics of the patient populations served.25-28 All sites that contributed data to this study were located within academic/teaching hospitals, which suggests that the variability was not due to hospital teaching status. Rates of highly resistant microorganisms were lowest in Manitoba and Quebec (5% each) and highest in Alberta, British Columbia and Ontario (20%, 16% and 15%, respectively). This suggests that the variability may be driven by geographical differences in hospital practices, patient populations or strain prevalence. Further investigation of this heterogeneity is warranted to determine whether there are effective strategies to reduce highly resistant microorganisms in highly prevalent settings. These findings also emphasize the importance of looking to local hospital antibiograms for guidance in selecting empiric treatment regimens.
Limitations
This study has limitations, given the secondary use of the data. The primary objective of the Bacteremia Antibiotic Length Actually Needed for Clinical Effectiveness study8 was to study duration of antibiotic treatment for bacteremia without deep-seated infection, and thus some infections were excluded (endocarditis, osteomyelitis, septic arthritis, undrained abscess and unremoved prosthetic material); however, these represented a small minority. In addition, the use of routine microbiological testing data rather than data expressly collected for the purpose of resistance surveillance means that not all organisms were tested for susceptibility to all potentially relevant agents and that there may have been variability in susceptibility testing methods across ICU/hospital sites. However, testing methods are generally standardized across Canada with the Clinical Laboratory Standards Institute guidelines.
Conclusion
We describe the epidemiologic features of bloodstream infections and corresponding antimicrobial susceptibility profiles in a Canadian multicentre cohort of critically ill patients determined with the use of routine microbiological testing data. Although this was a time-limited, single retrospective study, it provides proof-of-concept that routine susceptibility testing information from clinical isolates can be harnessed for antimicrobial resistance surveillance at the local level. In contrast to traditional microbiology surveillance networks, routine susceptibility testing offers the theoretical advantages of timeliness, efficiency of resources and ability to examine local trends. Consequently, we recommend that future data sharing could expand to a larger sample of ICUs across Canada, be conducted serially and have data analyzed and reported promptly; this would further improve the geographical representation and timeliness of these data. Such work is urgently needed to guide empiric treatment guidelines for patients as well as broader system-wide prevention methods to limit the spread of antimicrobial resistance.
Supplemental information
For reviewer comments and the original submission of this manuscript, please see www.cmajopen.ca/content/4/4/E569/suppl/DC1
Footnotes
Competing interests: None declared.
Contributors: Rachel Savage, Robert Fowler and Nick Daneman conceived and designed the study, developed the statistical analysis plan, analyzed and interpreted the data and drafted the manuscript. Asgar Rishu, Sean Bagshaw, Deborah Cook, Peter Dodek, Richard Hall, Anand Kumar, François Lamontagne, François Lauzier, John Marshall, Claudio Martin, Lauralyn McIntyre, John Muscedere, Steven Reynolds and Henry Stelfox participated in the study design and revised the manuscript for important intellectual content. All of the authors approved the final version to be published and agreed to act as guarantors of the work.
Disclaimer: This study was supported by the Institute for Clinical Evaluative Sciences (ICES), which is funded by an annual grant from the Ontario Ministry of Health and Long-Term Care (MOHLTC). The opinions, results and conclusions reported in this article are those of the authors and are independent from the funding sources. No endorsement by the ICES or the Ontario MOHLTC is intended or should be inferred.
References
- Copyright 2016, Joule Inc. or its licensors